Difference between revisions of "YrhJ"
| Line 15: | Line 15: | ||
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU27160 yrhJ] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU27160 yrhJ] | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/subtipathways/search.php?enzyme=YrhJ YrhJ]''' | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 118 kDa, 6.017 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 118 kDa, 6.017 | ||
Revision as of 11:48, 8 April 2014
- Description: cytochrome P450 (CYP102A3)/ NADPH-cytochrome P450 reductase
| Gene name | yrhJ |
| Synonyms | cypE |
| Essential | no |
| Product | NADPH-cytochrome P450 reductase |
| Function | fatty acid metabolism |
| Gene expression levels in SubtiExpress: yrhJ | |
| Metabolic function and regulation of this protein in SubtiPathways: YrhJ | |
| MW, pI | 118 kDa, 6.017 |
| Gene length, protein length | 3162 bp, 1054 aa |
| Immediate neighbours | rrcM, fatR |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 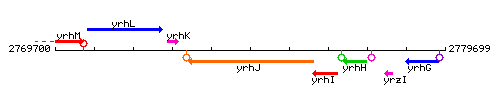
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed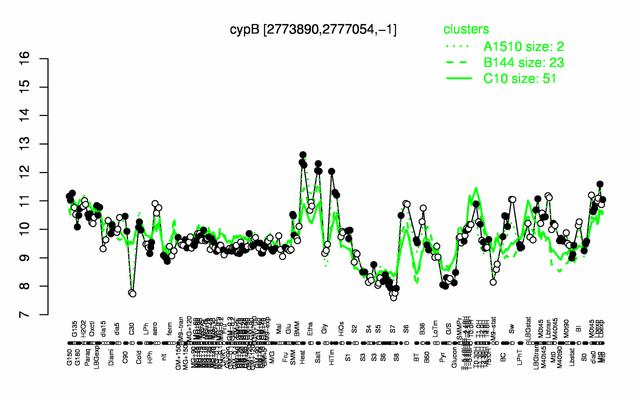
| |
Contents
Categories containing this gene/protein
electron transport/ other, lipid metabolism/ other, cell envelope stress proteins (controlled by SigM, V, W, X, Y), membrane proteins
This gene is a member of the following regulons
FatR regulon, SigM regulon, SigW regulon, SigX regulon
The gene
Basic information
- Locus tag: BSU27160
Phenotypes of a mutant
Database entries
- BsubCyc: BSU27160
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- hydroxylates medium-chain fatty acids in subterminal positions PubMed
- Protein family:
- Paralogous protein(s): YetO
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cell membrane (according to Swiss-Prot)
Database entries
- BsubCyc: BSU27160
- Structure: 2X7Y (from Bacillus megaterium; 67% identity, 88% similarity)
- UniProt: O08336
- KEGG entry: [3]
- E.C. number: 1.6.2.4
Additional information
Expression and regulation
- Regulation:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Veronica Guariglia-Oropeza, John D Helmann
Bacillus subtilis σ(V) confers lysozyme resistance by activation of two cell wall modification pathways, peptidoglycan O-acetylation and D-alanylation of teichoic acids.
J Bacteriol: 2011, 193(22);6223-32
[PubMed:21926231]
[WorldCat.org]
[DOI]
(I p)
Irnov Irnov, Cynthia M Sharma, Jörg Vogel, Wade C Winkler
Identification of regulatory RNAs in Bacillus subtilis.
Nucleic Acids Res: 2010, 38(19);6637-51
[PubMed:20525796]
[WorldCat.org]
[DOI]
(I p)
Sabine Eiben, Leonard Kaysser, Steffen Maurer, Katja Kühnel, Vlada B Urlacher, Rolf D Schmid
Preparative use of isolated CYP102 monooxygenases -- a critical appraisal.
J Biotechnol: 2006, 124(4);662-9
[PubMed:16716428]
[WorldCat.org]
[DOI]
(P p)
Oliver Lentz, Anton Feenstra, Tilo Habicher, Bernhard Hauer, Rolf D Schmid, Vlada B Urlacher
Altering the regioselectivity of cytochrome P450 CYP102A3 of Bacillus subtilis by using a new versatile assay system.
Chembiochem: 2006, 7(2);345-50
[PubMed:16381045]
[WorldCat.org]
[DOI]
(P p)
Mattias C U Gustafsson, Olivier Roitel, Ker R Marshall, Michael A Noble, Stephen K Chapman, Antonio Pessegueiro, Armand J Fulco, Myles R Cheesman, Claes von Wachenfeldt, Andrew W Munro
Expression, purification, and characterization of Bacillus subtilis cytochromes P450 CYP102A2 and CYP102A3: flavocytochrome homologues of P450 BM3 from Bacillus megaterium.
Biochemistry: 2004, 43(18);5474-87
[PubMed:15122913]
[WorldCat.org]
[DOI]
(P p)
Oliver Lentz, Vlada Urlacher, Rolf D Schmid
Substrate specificity of native and mutated cytochrome P450 (CYP102A3) from Bacillus subtilis.
J Biotechnol: 2004, 108(1);41-9
[PubMed:14741768]
[WorldCat.org]
[DOI]
(P p)
Penny D Thackray, Anne Moir
SigM, an extracytoplasmic function sigma factor of Bacillus subtilis, is activated in response to cell wall antibiotics, ethanol, heat, acid, and superoxide stress.
J Bacteriol: 2003, 185(12);3491-8
[PubMed:12775685]
[WorldCat.org]
[DOI]
(P p)
Min Cao, Tao Wang, Rick Ye, John D Helmann
Antibiotics that inhibit cell wall biosynthesis induce expression of the Bacillus subtilis sigma(W) and sigma(M) regulons.
Mol Microbiol: 2002, 45(5);1267-76
[PubMed:12207695]
[WorldCat.org]
[DOI]
(P p)
M C Gustafsson, C N Palmer, C R Wolf, C von Wachenfeldt
Fatty-acid-displaced transcriptional repressor, a conserved regulator of cytochrome P450 102 transcription in Bacillus species.
Arch Microbiol: 2001, 176(6);459-64
[PubMed:11734890]
[WorldCat.org]
[DOI]
(P p)
T R Lee, H P Hsu, G C Shaw
Transcriptional regulation of the Bacillus subtilis bscR-CYP102A3 operon by the BscR repressor and differential induction of cytochrome CYP102A3 expression by oleic acid and palmitate.
J Biochem: 2001, 130(4);569-74
[PubMed:11574077]
[WorldCat.org]
[DOI]
(P p)
C N Palmer, M C Gustafsson, H Dobson, C von Wachenfeldt, C R Wolf
Adaptive responses to fatty acids are mediated by the regulated expression of cytochromes P450.
Biochem Soc Trans: 1999, 27(4);374-8
[PubMed:10917605]
[WorldCat.org]
[DOI]
(P p)
X Huang, J D Helmann
Identification of target promoters for the Bacillus subtilis sigma X factor using a consensus-directed search.
J Mol Biol: 1998, 279(1);165-73
[PubMed:9636707]
[WorldCat.org]
[DOI]
(P p)