Difference between revisions of "SigB"
| Line 15: | Line 15: | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU04730 sigB] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU04730 sigB] | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=SigB SigB] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/stress_response.html Stress], [http://subtiwiki.uni-goettingen.de/pathways/murein/index.html Murein recycling]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/stress_response.html Stress], [http://subtiwiki.uni-goettingen.de/pathways/murein/index.html Murein recycling]''' | ||
Revision as of 15:15, 11 November 2013
- Description: RNA polymerase sigma factor SigB
| Gene name | sigB |
| Synonyms | rpoF |
| Essential | no |
| Product | RNA polymerase sigma factor SigB |
| Function | general stress response |
| Gene expression levels in SubtiExpress: sigB | |
| Interactions involving this protein in SubtInteract: SigB | |
| Metabolic function and regulation of this protein in SubtiPathways: Stress, Murein recycling | |
| MW, pI | 29 kDa, 5.418 |
| Gene length, protein length | 792 bp, 264 aa |
| Immediate neighbours | rsbW, rsbX |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 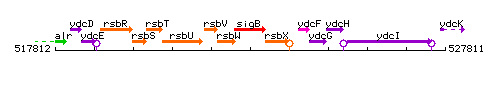
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed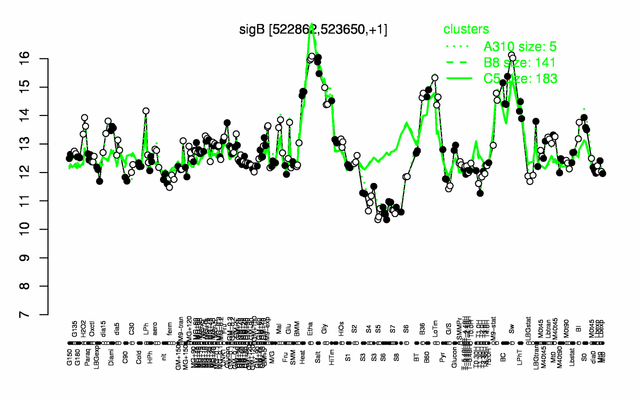
| |
Contents
Categories containing this gene/protein
transcription, sigma factors and their control, general stress proteins (controlled by SigB)
This gene is a member of the following regulons
The SigB regulon
The gene
Basic information
- Locus tag: BSU04730
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: SigB subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P06574
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant: QB5344 (cat), available in the Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Bill Haldenwang, San Antonio, USA
- Chet Price, Davis, USA homepage
Your additional remarks
References
Reviews
Control of SigB activity by protein-protein interactions
Oleg A Igoshin, Margaret S Brody, Chester W Price, Michael A Savageau
Distinctive topologies of partner-switching signaling networks correlate with their physiological roles.
J Mol Biol: 2007, 369(5);1333-52
[PubMed:17498739]
[WorldCat.org]
[DOI]
(P p)
Tae-Jong Kim, Tatiana A Gaidenko, Chester W Price
In vivo phosphorylation of partner switching regulators correlates with stress transmission in the environmental signaling pathway of Bacillus subtilis.
J Bacteriol: 2004, 186(18);6124-32
[PubMed:15342582]
[WorldCat.org]
[DOI]
(P p)
Tae-Jong Kim, Tatiana A Gaidenko, Chester W Price
A multicomponent protein complex mediates environmental stress signaling in Bacillus subtilis.
J Mol Biol: 2004, 341(1);135-50
[PubMed:15312768]
[WorldCat.org]
[DOI]
(P p)
Olivier Delumeau, Richard J Lewis, Michael D Yudkin
Protein-protein interactions that regulate the energy stress activation of sigma(B) in Bacillus subtilis.
J Bacteriol: 2002, 184(20);5583-9
[PubMed:12270815]
[WorldCat.org]
[DOI]
(P p)
M S Brody, K Vijay, C W Price
Catalytic function of an alpha/beta hydrolase is required for energy stress activation of the sigma(B) transcription factor in Bacillus subtilis.
J Bacteriol: 2001, 183(21);6422-8
[PubMed:11591687]
[WorldCat.org]
[DOI]
(P p)
C Eymann, M Hecker
Induction of sigma(B)-dependent general stress genes by amino acid starvation in a spo0H mutant of Bacillus subtilis.
FEMS Microbiol Lett: 2001, 199(2);221-7
[PubMed:11377871]
[WorldCat.org]
[DOI]
(P p)
A Dufour, U Voelker, A Voelker, W G Haldenwang
Relative levels and fractionation properties of Bacillus subtilis σ(B) and its regulators during balanced growth and stress.
J Bacteriol: 1996, 178(13);3701-9 sigma
[PubMed:8682769]
[WorldCat.org]
[DOI]
(P p)
U Voelker, A Voelker, B Maul, M Hecker, A Dufour, W G Haldenwang
Separate mechanisms activate sigma B of Bacillus subtilis in response to environmental and metabolic stresses.
J Bacteriol: 1995, 177(13);3771-80
[PubMed:7601843]
[WorldCat.org]
[DOI]
(P p)
A A Wise, C W Price
Four additional genes in the sigB operon of Bacillus subtilis that control activity of the general stress factor sigma B in response to environmental signals.
J Bacteriol: 1995, 177(1);123-33
[PubMed:8002610]
[WorldCat.org]
[DOI]
(P p)
A K Benson, W G Haldenwang
Bacillus subtilis sigma B is regulated by a binding protein (RsbW) that blocks its association with core RNA polymerase.
Proc Natl Acad Sci U S A: 1993, 90(6);2330-4
[PubMed:8460143]
[WorldCat.org]
[DOI]
(P p)
Identification of the SigB regulon
Priyanka Nannapaneni, Falk Hertwig, Maren Depke, Michael Hecker, Ulrike Mäder, Uwe Völker, Leif Steil, Sacha A F T van Hijum
Defining the structure of the general stress regulon of Bacillus subtilis using targeted microarray analysis and random forest classification.
Microbiology (Reading): 2012, 158(Pt 3);696-707
[PubMed:22174379]
[WorldCat.org]
[DOI]
(I p)
J D Helmann, M F Wu, P A Kobel, F J Gamo, M Wilson, M M Morshedi, M Navre, C Paddon
Global transcriptional response of Bacillus subtilis to heat shock.
J Bacteriol: 2001, 183(24);7318-28
[PubMed:11717291]
[WorldCat.org]
[DOI]
(P p)
A Petersohn, M Brigulla, S Haas, J D Hoheisel, U Völker, M Hecker
Global analysis of the general stress response of Bacillus subtilis.
J Bacteriol: 2001, 183(19);5617-31
[PubMed:11544224]
[WorldCat.org]
[DOI]
(P p)
C W Price, P Fawcett, H Cérémonie, N Su, C K Murphy, P Youngman
Genome-wide analysis of the general stress response in Bacillus subtilis.
Mol Microbiol: 2001, 41(4);757-74
[PubMed:11532142]
[WorldCat.org]
[DOI]
(P p)
A Petersohn, J Bernhardt, U Gerth, D Höper, T Koburger, U Völker, M Hecker
Identification of sigma(B)-dependent genes in Bacillus subtilis using a promoter consensus-directed search and oligonucleotide hybridization.
J Bacteriol: 1999, 181(18);5718-24
[PubMed:10482513]
[WorldCat.org]
[DOI]
(P p)
Anja Petersohn, Haike Antelmann, Ulf Gerth, Michael Hecker
Identification and transcriptional analysis of new members of the sigmaB regulon in Bacillus subtilis.
Microbiology (Reading): 1999, 145 ( Pt 4);869-880
[PubMed:10220166]
[WorldCat.org]
[DOI]
(P p)
Other publications