Difference between revisions of "PrkC"
| Line 29: | Line 29: | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| − | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=prkC_1651142_1653088_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:prkC_expression.png|500px]] | + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=prkC_1651142_1653088_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:prkC_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU15770]] |
|- | |- | ||
|} | |} | ||
Revision as of 13:19, 16 May 2013
- Description: protein kinase C, induce germination of spores in response to DAP-type, and not to Lys-type cell wall muropeptides
| Gene name | prkC |
| Synonyms | yloP |
| Essential | no |
| Product | protein kinase |
| Function | germination in response to muropeptides |
| Gene expression levels in SubtiExpress: prkC | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 71 kDa, 4.833 |
| Gene length, protein length | 1944 bp, 648 aa |
| Immediate neighbours | prpC, cpgA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 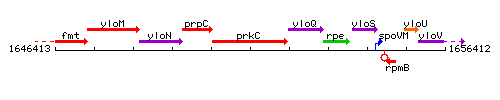
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed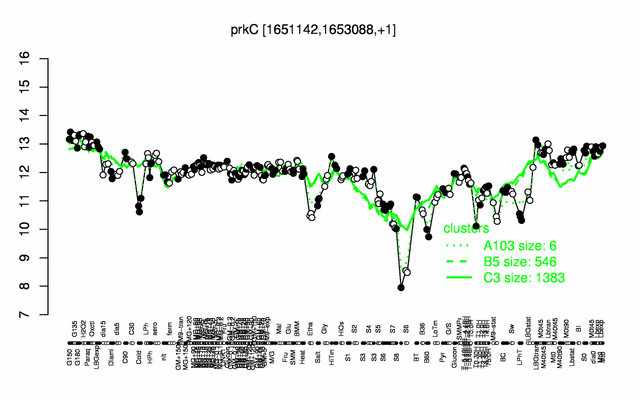
| |
Contents
Categories containing this gene/protein
protein modification, germination, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15770
Phenotypes of a mutant
- unable to germinate in response to muropeptides PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + a protein = ADP + a phosphoprotein (according to Swiss-Prot)
- Protein family: protein kinase domain (according to Swiss-Prot)
- Paralogous protein(s):
Proteins phosphorylated by PrkC
CpgA, EF-Tu, YezB PubMed, EF-G PubMed, YwjH, GlnA, Icd, AlsD, HPr PubMed
Extended information on the protein
- Kinetic information:
- Domains: PASTA domain at the C-terminus (binds muropeptides) PubMed
- Modification: phosphorylation on Thr-290 PubMed, autophosphorylation on multiple threonine residues PubMed
- Cofactor(s):
- Effectors of protein activity: activated by muropeptides PubMed
- Localization: inner spore membrane PubMed, membrane PubMed
Database entries
- UniProt: O34507
- KEGG entry: [2]
- E.C. number: 2.7.11.1
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP576 (spc), OMG302 (aphA3), available in Stülke lab
- Expression vector:
- for expression/ purification from B. subtilis with N-terminal Strep-tag, for SPINE, in pGP380: pGP832, available in Stülke lab
- for expression/ purification of the kinase domain from B. subtilis with N-terminal Strep-tag, for SPINE, in pGP380: pGP849, available in Stülke lab
- for expression, purification in E. coli with N-terminal His-tag, in pWH844: pGP1001, available in Stülke lab
- for expression, purification in E. coli with N-terminal Strep-tag, in pGP172: pGP825, available in Stülke lab
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Sandro F F Pereira, Lindsie Goss, Jonathan Dworkin
Eukaryote-like serine/threonine kinases and phosphatases in bacteria.
Microbiol Mol Biol Rev: 2011, 75(1);192-212
[PubMed:21372323]
[WorldCat.org]
[DOI]
(I p)
Jonathan Dworkin, Ishita M Shah
Exit from dormancy in microbial organisms.
Nat Rev Microbiol: 2010, 8(12);890-6
[PubMed:20972452]
[WorldCat.org]
[DOI]
(I p)
Phosphorylation of PrkC
Additional publications: PubMed
Targets of PrkC-dependent phosphorylation
Phsiological role of PrkC
Expression of PrkC: PubMed
Structure/ biochemistry of PrkC
Flavia Squeglia, Roberta Marchetti, Alessia Ruggiero, Rosa Lanzetta, Daniela Marasco, Jonathan Dworkin, Maxim Petoukhov, Antonio Molinaro, Rita Berisio, Alba Silipo
Chemical basis of peptidoglycan discrimination by PrkC, a key kinase involved in bacterial resuscitation from dormancy.
J Am Chem Soc: 2011, 133(51);20676-9
[PubMed:22111897]
[WorldCat.org]
[DOI]
(I p)
Alessia Ruggiero, Flavia Squeglia, Daniela Marasco, Roberta Marchetti, Antonio Molinaro, Rita Berisio
X-ray structural studies of the entire extracellular region of the serine/threonine kinase PrkC from Staphylococcus aureus.
Biochem J: 2011, 435(1);33-41
[PubMed:21208192]
[WorldCat.org]
[DOI]
(I p)