Difference between revisions of "MtnB"
| Line 14: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || methionine salvage | |style="background:#ABCDEF;" align="center"|'''Function''' || methionine salvage | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU13610 mtnB] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/cys_meth_and_sulfate_assimilation.html Cys, Met & Sulfate assimilation]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/cys_meth_and_sulfate_assimilation.html Cys, Met & Sulfate assimilation]''' | ||
| Line 24: | Line 24: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[mtnX]]'', ''[[mtnD]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[mtnX]]'', ''[[mtnD]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU13610 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU13610 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU13610 Advanced_DNA] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:ykrY_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:ykrY_context.gif]] | ||
Revision as of 12:40, 13 May 2013
- Description: methylthioribulose-1-phosphate dehydratase
| Gene name | mtnB |
| Synonyms | ykrY |
| Essential | no |
| Product | methylthioribulose-1-phosphate dehydratase |
| Function | methionine salvage |
| Gene expression levels in SubtiExpress: mtnB | |
| Metabolic function and regulation of this protein in SubtiPathways: Cys, Met & Sulfate assimilation | |
| MW, pI | 23 kDa, 5.686 |
| Gene length, protein length | 627 bp, 209 aa |
| Immediate neighbours | mtnX, mtnD |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 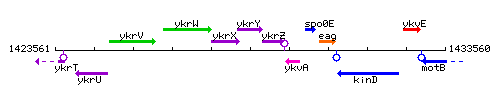
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed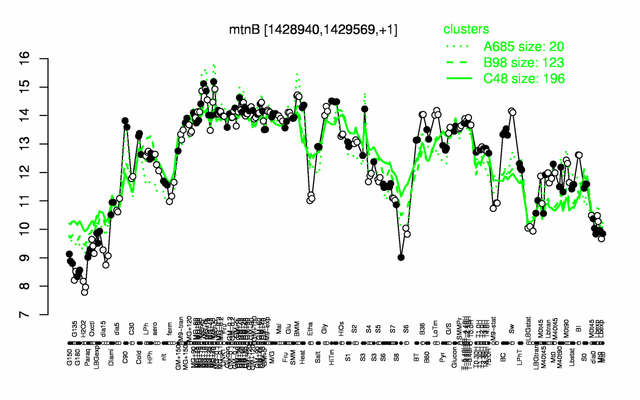
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13610
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: S-methyl-5-thio-D-ribulose 1-phosphate = 5-(methylthio)-2,3-dioxopentyl phosphate + H2O (according to Swiss-Prot)
- Protein family: MtnB subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O31668
- KEGG entry: [3]
- E.C. number: 4.2.1.109
Additional information
Expression and regulation
- Regulatory mechanism: S-box: transcription termination/ antitermination, the S-box riboswitch binds S-adenosylmethionine resulting in termination PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Jerneja Tomsic, Brooke A McDaniel, Frank J Grundy, Tina M Henkin
Natural variability in S-adenosylmethionine (SAM)-dependent riboswitches: S-box elements in bacillus subtilis exhibit differential sensitivity to SAM In vivo and in vitro.
J Bacteriol: 2008, 190(3);823-33
[PubMed:18039762]
[WorldCat.org]
[DOI]
(I p)
Agnieszka Sekowska, Valérie Dénervaud, Hiroki Ashida, Karine Michoud, Dieter Haas, Akiho Yokota, Antoine Danchin
Bacterial variations on the methionine salvage pathway.
BMC Microbiol: 2004, 4;9
[PubMed:15102328]
[WorldCat.org]
[DOI]
(I e)
Maumita Mandal, Benjamin Boese, Jeffrey E Barrick, Wade C Winkler, Ronald R Breaker
Riboswitches control fundamental biochemical pathways in Bacillus subtilis and other bacteria.
Cell: 2003, 113(5);577-86
[PubMed:12787499]
[WorldCat.org]
[DOI]
(P p)
Ulrike Mäder, Georg Homuth, Christian Scharf, Knut Büttner, Rüdiger Bode, Michael Hecker
Transcriptome and proteome analysis of Bacillus subtilis gene expression modulated by amino acid availability.
J Bacteriol: 2002, 184(15);4288-95
[PubMed:12107147]
[WorldCat.org]
[DOI]
(P p)
Agnieszka Sekowska, Antoine Danchin
The methionine salvage pathway in Bacillus subtilis.
BMC Microbiol: 2002, 2;8
[PubMed:12022921]
[WorldCat.org]
[DOI]
(I e)
Brooke A Murphy, Frank J Grundy, Tina M Henkin
Prediction of gene function in methylthioadenosine recycling from regulatory signals.
J Bacteriol: 2002, 184(8);2314-8
[PubMed:11914366]
[WorldCat.org]
[DOI]
(P p)