Difference between revisions of "YodB"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
|||
| Line 57: | Line 57: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
=The protein= | =The protein= | ||
Revision as of 19:14, 27 January 2012
- Description: MarR/DUF24 family transcription repressor of azoR1, catD-catE and yodC expression
| Gene name | yodB |
| Synonyms | |
| Essential | no |
| Product | MarR/DUF24 family transcription repressor |
| Function | regulation of quinone and diamide detoxification |
| MW, pI | 12 kDa, 4.737 |
| Gene length, protein length | 336 bp, 112 aa |
| Immediate neighbours | yodA, yodC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 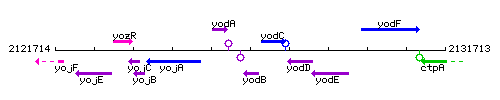
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
transcription factors and their control, resistance against oxidative and electrophile stress
This gene is a member of the following regulons
The YodB regulon:
The gene
Basic information
- Locus tag: BSU19540
Phenotypes of a mutant
- increased resistance to catechol, quinones and diamide PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: MarR/DUF24 family
- Paralogous protein(s): CatR
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: redox-controlled by intersubunit disulfide formation of Cys6 and Cys101' in response to quinones and diamide
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O34844
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulation:
- subject to autorepression PubMed
- Regulatory mechanism:
- YodB: transcription repression PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Haike Antelmann,University of Greifswald, Germany
Your additional remarks
References
Reviews
Antelmann H, Helmann JD. Thiol-based redox switches and gene regulation. Antioxid Redox Signal. 2011,14:1049-63. PubMed
Original Publications