Difference between revisions of "YjbC"
(→Categories containing this gene/protein) |
|||
| Line 35: | Line 35: | ||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
{{SubtiWiki category|[[general stress proteins (controlled by SigB)]]}}, | {{SubtiWiki category|[[general stress proteins (controlled by SigB)]]}}, | ||
| − | {{SubtiWiki category|[[cell envelope stress proteins (controlled by SigM, W, X, Y)]]}} | + | {{SubtiWiki category|[[cell envelope stress proteins (controlled by SigM, V, W, X, Y)]]}} |
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
Revision as of 06:15, 25 August 2011
- Description: general stress protein, required for survival of salt stress
| Gene name | yjbC |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | unknown |
| MW, pI | 22 kDa, 5.14 |
| Gene length, protein length | 576 bp, 192 aa |
| Immediate neighbours | yjbB, spx |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 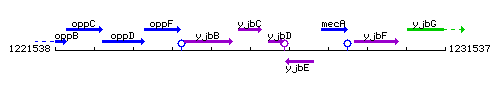
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
general stress proteins (controlled by SigB), cell envelope stress proteins (controlled by SigM, V, W, X, Y)
This gene is a member of the following regulons
PerR regulon, SigB regulon, SigM regulon, SigW regulon, SigX regulon
The gene
Basic information
- Locus tag: BSU11490
Phenotypes of a mutant
strongly impaired survival of salt stress PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- UniProt: O31601
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Montira Leelakriangsak, Kazuo Kobayashi, Peter Zuber
Dual negative control of spx transcription initiation from the P3 promoter by repressors PerR and YodB in Bacillus subtilis.
J Bacteriol: 2007, 189(5);1736-44
[PubMed:17158660]
[WorldCat.org]
[DOI]
(P p)
Dirk Höper, Uwe Völker, Michael Hecker
Comprehensive characterization of the contribution of individual SigB-dependent general stress genes to stress resistance of Bacillus subtilis.
J Bacteriol: 2005, 187(8);2810-26
[PubMed:15805528]
[WorldCat.org]
[DOI]
(P p)
H Antelmann, C Scharf, M Hecker
Phosphate starvation-inducible proteins of Bacillus subtilis: proteomics and transcriptional analysis.
J Bacteriol: 2000, 182(16);4478-90
[PubMed:10913081]
[WorldCat.org]
[DOI]
(P p)