Difference between revisions of "SigB"
| Line 84: | Line 84: | ||
* '''Structure:''' | * '''Structure:''' | ||
| − | * ''' | + | * '''UniProt:''' [http://www.uniprot.org/uniprot/P06574 P06574] |
* '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU04730] | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU04730] | ||
Revision as of 14:37, 21 July 2009
- Description: RNA polymerase sigma factor SigB
| Gene name | sigB |
| Synonyms | rpoF |
| Essential | no |
| Product | RNA polymerase sigma factor SigB |
| Function | general stress response |
| Metabolic function and regulation of this protein in SubtiPathways: Stress | |
| MW, pI | 29 kDa, 5.418 |
| Gene length, protein length | 792 bp, 264 aa |
| Immediate neighbours | rsbW, rsbX |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 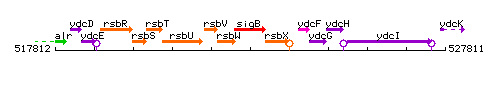
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU04730
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: SigB subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Genes/ operons controlled by SigB
csbB, ctc, gsiB, katE, lexA,licR, sfA, xpf, yaaH, ydeC, ytzE, yvaN, ywhH,
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- Structure:
- UniProt: P06574
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Bill Haldenwang, San Antonio, USA
- Chet Price, Davis, USA homepage
Your additional remarks
References
Reviews
Peter Zuber
Management of oxidative stress in Bacillus.
Annu Rev Microbiol: 2009, 63;575-97
[PubMed:19575568]
[WorldCat.org]
[DOI]
(I p)
Michael Hecker, Jan Pané-Farré, Uwe Völker
SigB-dependent general stress response in Bacillus subtilis and related gram-positive bacteria.
Annu Rev Microbiol: 2007, 61;215-36
[PubMed:18035607]
[WorldCat.org]
[DOI]
(P p)
M Hecker, U Völker
General stress response of Bacillus subtilis and other bacteria.
Adv Microb Physiol: 2001, 44;35-91
[PubMed:11407115]
[WorldCat.org]
[DOI]
(P p)
M Hecker, U Völker
Non-specific, general and multiple stress resistance of growth-restricted Bacillus subtilis cells by the expression of the sigmaB regulon.
Mol Microbiol: 1998, 29(5);1129-36
[PubMed:9767581]
[WorldCat.org]
[DOI]
(P p)
Control of SigB activity by protein-protein interactions
Identification of the SigB regulon
J D Helmann, M F Wu, P A Kobel, F J Gamo, M Wilson, M M Morshedi, M Navre, C Paddon
Global transcriptional response of Bacillus subtilis to heat shock.
J Bacteriol: 2001, 183(24);7318-28
[PubMed:11717291]
[WorldCat.org]
[DOI]
(P p)
A Petersohn, M Brigulla, S Haas, J D Hoheisel, U Völker, M Hecker
Global analysis of the general stress response of Bacillus subtilis.
J Bacteriol: 2001, 183(19);5617-31
[PubMed:11544224]
[WorldCat.org]
[DOI]
(P p)
C W Price, P Fawcett, H Cérémonie, N Su, C K Murphy, P Youngman
Genome-wide analysis of the general stress response in Bacillus subtilis.
Mol Microbiol: 2001, 41(4);757-74
[PubMed:11532142]
[WorldCat.org]
[DOI]
(P p)
A Petersohn, J Bernhardt, U Gerth, D Höper, T Koburger, U Völker, M Hecker
Identification of sigma(B)-dependent genes in Bacillus subtilis using a promoter consensus-directed search and oligonucleotide hybridization.
J Bacteriol: 1999, 181(18);5718-24
[PubMed:10482513]
[WorldCat.org]
[DOI]
(P p)
Anja Petersohn, Haike Antelmann, Ulf Gerth, Michael Hecker
Identification and transcriptional analysis of new members of the sigmaB regulon in Bacillus subtilis.
Microbiology (Reading): 1999, 145 ( Pt 4);869-880
[PubMed:10220166]
[WorldCat.org]
[DOI]
(P p)
Other publications
Ewelina Młynarska, Ewa Radzioch, Bartłomiej Dąbek, Klaudia Leszto, Alicja Witkowska, Witold Czarnik, Weronika Jędraszak, Jacek Rysz, Beata Franczyk
Hypertrophic Cardiomyopathy with Special Focus on Mavacamten and Its Future in Cardiology.
Biomedicines: 2024, 12(12);
[PubMed:39767581]
[WorldCat.org]
[DOI]
(P e)
Adam Reeves, Ulf Gerth, Uwe Völker, W G Haldenwang
ClpP modulates the activity of the Bacillus subtilis stress response transcription factor, sigmaB.
J Bacteriol: 2007, 189(17);6168-75
[PubMed:17586624]
[WorldCat.org]
[DOI]
(P p)
Noriko Suzuki, Naoki Takaya, Takayuki Hoshino, Akira Nakamura
Enhancement of a sigma(B)-dependent stress response in Bacillus subtilis by light via YtvA photoreceptor.
J Gen Appl Microbiol: 2007, 53(2);81-8
[PubMed:17575448]
[WorldCat.org]
[DOI]
(P p)
Dirk Höper, Uwe Völker, Michael Hecker
Comprehensive characterization of the contribution of individual SigB-dependent general stress genes to stress resistance of Bacillus subtilis.
J Bacteriol: 2005, 187(8);2810-26
[PubMed:15805528]
[WorldCat.org]
[DOI]
(P p)
Gudrun Holtmann, Matthias Brigulla, Leif Steil, Alexandra Schütz, Karsta Barnekow, Uwe Völker, Erhard Bremer
RsbV-independent induction of the SigB-dependent general stress regulon of Bacillus subtilis during growth at high temperature.
J Bacteriol: 2004, 186(18);6150-8
[PubMed:15342585]
[WorldCat.org]
[DOI]
(P p)
Karen Carniol, Tae-Jong Kim, Chester W Price, Richard Losick
Insulation of the sigmaF regulatory system in Bacillus subtilis.
J Bacteriol: 2004, 186(13);4390-4
[PubMed:15205443]
[WorldCat.org]
[DOI]
(P p)
Thorsten Mascher, Neil G Margulis, Tao Wang, Rick W Ye, John D Helmann
Cell wall stress responses in Bacillus subtilis: the regulatory network of the bacitracin stimulon.
Mol Microbiol: 2003, 50(5);1591-604
[PubMed:14651641]
[WorldCat.org]
[DOI]
(P p)
Shuyu Zhang, W G Haldenwang
RelA is a component of the nutritional stress activation pathway of the Bacillus subtilis transcription factor sigma B.
J Bacteriol: 2003, 185(19);5714-21
[PubMed:13129942]
[WorldCat.org]
[DOI]
(P p)
Matthias Brigulla, Tamara Hoffmann, Andrea Krisp, Andrea Völker, Erhard Bremer, Uwe Völker
Chill induction of the SigB-dependent general stress response in Bacillus subtilis and its contribution to low-temperature adaptation.
J Bacteriol: 2003, 185(15);4305-14
[PubMed:12867438]
[WorldCat.org]
[DOI]
(P p)
Claudia Rollenhagen, Haike Antelmann, Janine Kirstein, Olivier Delumeau, Michael Hecker, Michael D Yudkin
Binding of sigma(A) and sigma(B) to core RNA polymerase after environmental stress in Bacillus subtilis.
J Bacteriol: 2003, 185(1);35-40
[PubMed:12486038]
[WorldCat.org]
[DOI]
(P p)
A Petersohn, S Engelmann, P Setlow, M Hecker
The katX gene of Bacillus subtilis is under dual control of sigmaB and sigmaF.
Mol Gen Genet: 1999, 262(1);173-9
[PubMed:10503549]
[WorldCat.org]
[DOI]
(P p)
U Völker, B Maul, M Hecker
Expression of the sigmaB-dependent general stress regulon confers multiple stress resistance in Bacillus subtilis.
J Bacteriol: 1999, 181(13);3942-8
[PubMed:10383961]
[WorldCat.org]
[DOI]
(P p)
T Schweder, A Kolyschkow, U Völker, M Hecker
Analysis of the expression and function of the sigmaB-dependent general stress regulon of Bacillus subtilis during slow growth.
Arch Microbiol: 1999, 171(6);439-43
[PubMed:10369900]
[WorldCat.org]
[DOI]
(P p)
C von Blohn, B Kempf, R M Kappes, E Bremer
Osmostress response in Bacillus subtilis: characterization of a proline uptake system (OpuE) regulated by high osmolarity and the alternative transcription factor sigma B.
Mol Microbiol: 1997, 25(1);175-87
[PubMed:11902719]
[WorldCat.org]
[DOI]
(P p)
S Alper, A Dufour, D A Garsin, L Duncan, R Losick
Role of adenosine nucleotides in the regulation of a stress-response transcription factor in Bacillus subtilis.
J Mol Biol: 1996, 260(2);165-77
[PubMed:8764398]
[WorldCat.org]
[DOI]
(P p)
A R Redfield, C W Price
General stress transcription factor sigmaB of Bacillus subtilis is a stable protein.
J Bacteriol: 1996, 178(12);3668-70
[PubMed:8655572]
[WorldCat.org]
[DOI]
(P p)
S A Boylan, A R Redfield, M S Brody, C W Price
Stress-induced activation of the sigma B transcription factor of Bacillus subtilis.
J Bacteriol: 1993, 175(24);7931-7
[PubMed:8253681]
[WorldCat.org]
[DOI]
(P p)
A K Benson, W G Haldenwang
Regulation of sigma B levels and activity in Bacillus subtilis.
J Bacteriol: 1993, 175(8);2347-56
[PubMed:8468294]
[WorldCat.org]
[DOI]
(P p)
A K Benson, W G Haldenwang
The sigma B-dependent promoter of the Bacillus subtilis sigB operon is induced by heat shock.
J Bacteriol: 1993, 175(7);1929-35
[PubMed:8458834]
[WorldCat.org]
[DOI]
(P p)
C Ray, M Igo, W Shafer, R Losick, C P Moran
Suppression of ctc promoter mutations in Bacillus subtilis.
J Bacteriol: 1988, 170(2);900-7
[PubMed:3123466]
[WorldCat.org]
[DOI]
(P p)
M Igo, M Lampe, C Ray, W Schafer, C P Moran, R Losick
Genetic studies of a secondary RNA polymerase sigma factor in Bacillus subtilis.
J Bacteriol: 1987, 169(8);3464-9
[PubMed:3112122]
[WorldCat.org]
[DOI]
(P p)
M L Duncan, S S Kalman, S M Thomas, C W Price
Gene encoding the 37,000-dalton minor sigma factor of Bacillus subtilis RNA polymerase: isolation, nucleotide sequence, chromosomal locus, and cryptic function.
J Bacteriol: 1987, 169(2);771-8
[PubMed:3027048]
[WorldCat.org]
[DOI]
(P p)
M M Igo, R Losick
Regulation of a promoter that is utilized by minor forms of RNA polymerase holoenzyme in Bacillus subtilis.
J Mol Biol: 1986, 191(4);615-24
[PubMed:3100810]
[WorldCat.org]
[DOI]
(P p)
C Binnie, M Lampe, R Losick
Gene encoding the sigma 37 species of RNA polymerase sigma factor from Bacillus subtilis.
Proc Natl Acad Sci U S A: 1986, 83(16);5943-7
[PubMed:3016731]
[WorldCat.org]
[DOI]
(P p)
W C Johnson, C P Moran, R Losick
Two RNA polymerase sigma factors from Bacillus subtilis discriminate between overlapping promoters for a developmentally regulated gene.
Nature: 1983, 302(5911);800-4
[PubMed:6405278]
[WorldCat.org]
[DOI]
(P p)
J F Ollington, W G Haldenwang, T V Huynh, R Losick
Developmentally regulated transcription in a cloned segment of the Bacillus subtilis chromosome.
J Bacteriol: 1981, 147(2);432-42
[PubMed:6790515]
[WorldCat.org]
[DOI]
(P p)
W G Haldenwang, R Losick
Novel RNA polymerase sigma factor from Bacillus subtilis.
Proc Natl Acad Sci U S A: 1980, 77(12);7000-4
[PubMed:6784117]
[WorldCat.org]
[DOI]
(P p)
W G Haldenwang, R Losick
A modified RNA polymerase transcribes a cloned gene under sporulation control in Bacillus subtilis.
Nature: 1979, 282(5736);256-60
[PubMed:116131]
[WorldCat.org]
[DOI]
(P p)