Difference between revisions of "DnaN"
| Line 82: | Line 82: | ||
* '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/P05649 P05649] | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/P05649 P05649] | ||
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU00020] |
* '''E.C. number:''' [http://www.expasy.org/enzyme/2.7.7.7 2.7.7.7] | * '''E.C. number:''' [http://www.expasy.org/enzyme/2.7.7.7 2.7.7.7] | ||
Revision as of 02:01, 25 June 2009
- Description: DNA polymerase III (beta subunit), beta clamp
| Gene name | dnaN |
| Synonyms | dnaG, dnaK |
| Essential | yes PubMed |
| Product | DNA polymerase III (beta subunit), beta clamp |
| Function | DNA replication |
| MW, pI | 41 kDa, 4.718 |
| Gene length, protein length | 1134 bp, 378 aa |
| Immediate neighbours | dnaA, yaaA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 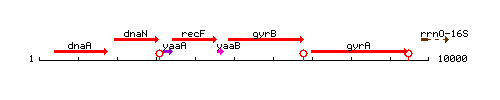
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU00020
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Deoxynucleoside triphosphate + DNA(n) = diphosphate + DNA(n+1) (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot), Nucleoid (Mid-cell) PubMed nucleoid (mid-cell spot) PubMed
Database entries
- Structure:
- Swiss prot entry: P05649
- KEGG entry: [3]
- E.C. number: 2.7.7.7
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Philippe Noirot, Jouy-en-Josas, France homepage
Your additional remarks
References
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
Marie-Françoise Noirot-Gros, M Velten, M Yoshimura, S McGovern, T Morimoto, S D Ehrlich, N Ogasawara, P Polard, Philippe Noirot
Functional dissection of YabA, a negative regulator of DNA replication initiation in Bacillus subtilis.
Proc Natl Acad Sci U S A: 2006, 103(7);2368-73
[PubMed:16461910]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)
Marie-Françoise Noirot-Gros, Etienne Dervyn, Ling Juan Wu, Peggy Mervelet, Jeffery Errington, S Dusko Ehrlich, Philippe Noirot
An expanded view of bacterial DNA replication.
Proc Natl Acad Sci U S A: 2002, 99(12);8342-7
[PubMed:12060778]
[WorldCat.org]
[DOI]
(P p)
T Fukuoka, S Moriya, H Yoshikawa, N Ogasawara
Purification and characterization of an initiation protein for chromosomal replication, DnaA, in Bacillus subtilis.
J Biochem: 1990, 107(5);732-9
[PubMed:2168872]
[WorldCat.org]
[DOI]
(P p)
N Ogasawara, S Moriya, H Yoshikawa
Structure and function of the region of the replication origin of the Bacillus subtilis chromosome. IV. Transcription of the oriC region and expression of DNA gyrase genes and other open reading frames.
Nucleic Acids Res: 1985, 13(7);2267-79
[PubMed:2987848]
[WorldCat.org]
[DOI]
(P p)